25. Linkage analysis of genes, ge (giant embryo) and esp-l
(endosperm storage protein-1) on chromosome 7
Toshihiro Kumamaru and Hikaru
Satoh
1) Plant Breeding and Genetic Research Laboratory, Japan Tobacco Inc.,
Higashibara, Toyoda. Iwata,
Shizuoka 438, Japan.
2) Faculty of Agriculture, Kyushu University, Hakozaki, Fukuoka 812,
Japan
Various kinds of mutants for endosperm traits have been
obtained by the treatment of fertilized egg cells with N-methyl-N-nitrosourea
(Satoh and Omura 1981; Satoh 1985). These mutants are useful for the improvement
of rice grain quality and also for linkage mapping. The trisomic analysis
for two mutant, ge for a giant embryo (Satoh and Omura 1981) and
esp-l
(endosperm storage protein mutant-1) characterizing the low content of
13b polypeptide (prolamin)(Kumamaru et al. 1988), indicated that
the two genes are located on chromosome 7 (Kumamaru et al. 1987;
Satoh and lwata 1990) and ge linked with v-ll and rfs,
with the recombination values of 38.9 ±4.1% and 20.9 ± 2.9%,
respectively (Satoh and lwata 1990). The detailed map of chromosome 7 showing
positions of these genes has not been prepared.
CM21 (esp-l) was crossed with EM40 (ge)
and some of marker lines with d-6, g, spl-5, v-ll and rfs,
which are located on chromosome 7. The F2 and F3 data show that esp-l
is linked tightly with ge and rfs with the recombination
values of 4.7 ± 1.5% and 20.6 ± 3.4%, respectively, and no
linkage was detected between esp-l, d-6, g, spl-5 and v-11
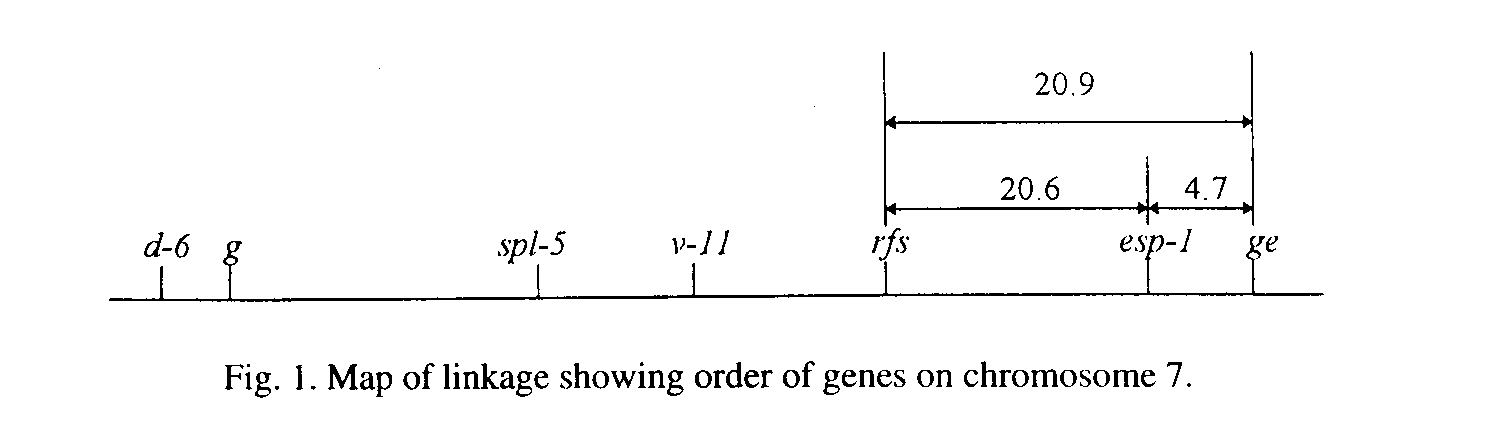
(Table 1 and 2). From these results, the order of the genes on chromosome
7 was estimated to be d-6� g�spl-5� v-ll�rfs-esp-l-ge as shown in
Fig. 1.
| Table 1. Linkage analysis
between esp-l and marker genes on chromosome 7 |
| Gene pair |
Phase |
Segregation
in F2 |
|
X2 |
| AB |
Ab |
aB |
ab |
Total |
(9:3:3:1) |
| esp-l X d-6 |
Repulsion |
178 |
52 |
47 |
17 |
294 |
0.44 |
| esp-I X g |
Repulsion |
186 |
49 |
50 |
13 |
298 |
0.01 |
| esp-I X spl-5 |
Repulsion |
179 |
58 |
57 |
11 |
305 |
1.64 |
| esp-l X v-ll |
Repulsion |
108 |
30 |
26 |
3 |
167 |
0.73 |
| esp-I Xspl-5 |
Repulsion |
84 |
51 |
37 |
4 |
176 |
13.1** |
**: Significant at 1% level.
Table 2. Segregation patterns in F3 derived from crosses
between
esp-l and two marker genes, ge and
rfs
|
+ |
|
|
esp-l |
|
|
| ++ |
+ge |
ge ge |
++ |
+ge |
ge ge |
R.V. |
| 8 |
99 |
50 |
42 |
1 |
0 |
4.7% ± 1.5 |
|
+ |
|
|
esp-l |
|
|
| ++ |
+rfs |
rfs rfs |
++ |
+rfs |
rfs rfs |
R.V. |
| 21 |
77 |
38 |
25 |
12 |
0 |
20.6%±3.4 |
 References
References
Kumamani, T., H. Satoh, N. Iwata, T. Omura and M. Ogawa, 1987. Mutants
for rice storage proteins. III.
Genetic analysis of mutants for storage proteins
of protein bodies in the starchy endosperm. Japan. J. Breed.
62:333-339.
Kumamani, T., H. Satoh, N. Iwata, T. Omura, M. Ogawa and K. Tanaka,
1987. Mutants for rice storage
proteins. 1. Screening of mutants for storage proteins
of protein bodies in the starchy endosperm.
Theor. Appl. Genet., 76:11-16.
Satoh, H. and T. Omura, 1979. New endosperm mutations induced by chemical
mutagens in rice, Oryza sativa
L. Japan. J. Breed. 31: 316-326.
Satoh, H. and N. Iwata, 1990. Linkage analysis in rice. On three mutant
loci for endosperm properties,
ge (giant embryo), du-4 (dull endosperm-4)
and flo-I (floury endospcrm-l). Japan J. Breed. 47
(Suppl. 2): 268-269. (in Japanese)
